RFP Lab
Purpose:
- Make RFP from jellyfish bacteria
- Learn about steps genetic engineering
- 2a- materials + procedure can be found in Amgen lab manual part 2a
- 4a- materials + procedure can be found in Amgen lab manual part 4a
- 5a- materials + procedure can be found in Amgen lab manual part 5a
- 6a- materials + procedure can be found in Amgen lab manual part 6a
| abridged_biotech_sequence_amgen_1.pdf | |
| File Size: | 3854 kb |
| File Type: | |
Experimental Overview:
Part 2a:
-Verification of plasmid by restriction digest
-Cut plasmid with BanH1 + Hind III to cut out RFP-ara from bacterial plasmid
Part 4a:
-Verification of plasmid digest by electrophoresis
Part 5a:
-Transformation of bacteria with recombinant plasmid
Part 6:
-Purification of RFP using chromatography
Part 2a:
-Verification of plasmid by restriction digest
-Cut plasmid with BanH1 + Hind III to cut out RFP-ara from bacterial plasmid
Part 4a:
-Verification of plasmid digest by electrophoresis
Part 5a:
-Transformation of bacteria with recombinant plasmid
Part 6:
-Purification of RFP using chromatography
Data:
2a - 1. ori-origin of replication, GoI-gene of interest, Amp-R-selective marker, Ara-C-binds to the promoter so we can get transcription of our GoI. 2. restriction enzymes in nature are defense mechanisms that cut DNA from other bacteria. 3. Bacteria retains an antibiotic resistance because otherwise they would die off. In the medical field they have to create a whole new class of antibiotics. 4. Bacteria have DNA, RNA polymerase, and ribosomes (central dogma) just like humans. We can replace DNA with other DNA that can change a specific trait.
4a - 1. Since the recombinant plasmid has the desired trait you want to make sure that it really has the desired trait. 2. The gel results compared different to our comparisons because the DNA ladder was not visible due to a malfunction somewhere. 3. No I don't believe so, besides the opposite of not showing up. 4. Well without the DNA ladder we can't conclude this question as for evidence because we cannot estimate the measures. 5. No just one band. 6. RFP to R+ and AMP R to R-.
5a - 1. My predictions were correct but we did miss one of our pipeting instances where we crossed the middle line. 2. None. 3. They did not appear.4. because not a lot is used as desirable, the more the better. 5. RFP gives the red flourescent glow to our proteins. 6. bacteria can make human protein because they have like organelles and cell processes.
2a - 1. ori-origin of replication, GoI-gene of interest, Amp-R-selective marker, Ara-C-binds to the promoter so we can get transcription of our GoI. 2. restriction enzymes in nature are defense mechanisms that cut DNA from other bacteria. 3. Bacteria retains an antibiotic resistance because otherwise they would die off. In the medical field they have to create a whole new class of antibiotics. 4. Bacteria have DNA, RNA polymerase, and ribosomes (central dogma) just like humans. We can replace DNA with other DNA that can change a specific trait.
4a - 1. Since the recombinant plasmid has the desired trait you want to make sure that it really has the desired trait. 2. The gel results compared different to our comparisons because the DNA ladder was not visible due to a malfunction somewhere. 3. No I don't believe so, besides the opposite of not showing up. 4. Well without the DNA ladder we can't conclude this question as for evidence because we cannot estimate the measures. 5. No just one band. 6. RFP to R+ and AMP R to R-.
5a - 1. My predictions were correct but we did miss one of our pipeting instances where we crossed the middle line. 2. None. 3. They did not appear.4. because not a lot is used as desirable, the more the better. 5. RFP gives the red flourescent glow to our proteins. 6. bacteria can make human protein because they have like organelles and cell processes.
Conclusion:
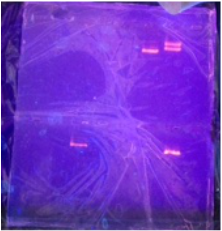
In the end, our red fluorescent protein did not show up, it turns out that the gene we received did not work properly so our end result wasn't how we expected it to be. After doing the lab again. The protein was pure because it appeared to be only one DNA band. The protein wasn't extremely concentrated since it wasn't very bold but was still visible. Our protein was visible around the 15-25 mark on the DNA ladder. Our protein matched fairly well to the desired protein level on the DNA ladder. This means our whole lab worked even though it may be hard to see.
Reflection:
This lab definitely was a long and attention to detail lab. It made us focus on every step ensuring we did it correctly and followed instructions. My group collaborated very well and was on task throughout the project. We were sure that we followed the guidelines and kept everything organized and structured. Even though our conclusion didn't come out as expected, we were still content on how we preformed the lab. We could could have asked more questions when we were not sure of the answer or the correct step and we could have been faster with our transitions from one procedure to another however we still finished fairly fast. Overall, this was a very interesting lab, even though we didn't end up with visible red fluorescent bacteria.
In the end, our red fluorescent protein did not show up, it turns out that the gene we received did not work properly so our end result wasn't how we expected it to be. After doing the lab again. The protein was pure because it appeared to be only one DNA band. The protein wasn't extremely concentrated since it wasn't very bold but was still visible. Our protein was visible around the 15-25 mark on the DNA ladder. Our protein matched fairly well to the desired protein level on the DNA ladder. This means our whole lab worked even though it may be hard to see.
Reflection:
This lab definitely was a long and attention to detail lab. It made us focus on every step ensuring we did it correctly and followed instructions. My group collaborated very well and was on task throughout the project. We were sure that we followed the guidelines and kept everything organized and structured. Even though our conclusion didn't come out as expected, we were still content on how we preformed the lab. We could could have asked more questions when we were not sure of the answer or the correct step and we could have been faster with our transitions from one procedure to another however we still finished fairly fast. Overall, this was a very interesting lab, even though we didn't end up with visible red fluorescent bacteria.